For optimal use, please specify the intended use for this tracer (receptor binding, RIA, etc.) in the order notes field before checking out.
Catalog # |
Size |
Price |
|
|---|---|---|---|
| T-075-40 | 10 µCi | $1167 |
 )
)
|
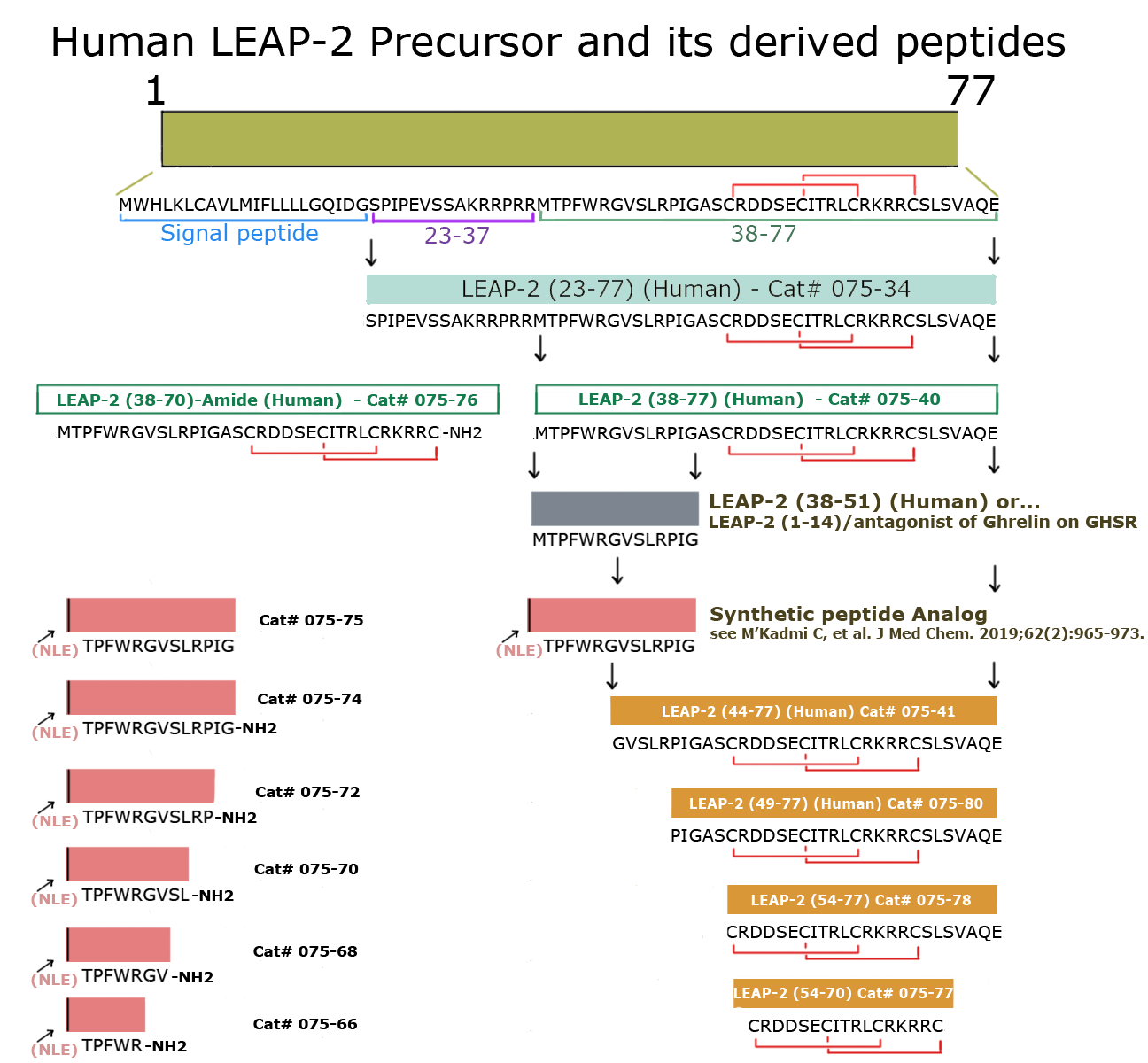
Tyr-Met-Thr-Pro-Phe-Trp-Arg-Gly-Val-Ser-Leu-Arg-Pro-Ile-Gly-Ala-Ser-Cys-Arg-Asp-Asp-Ser-Glu-Cys-Ile-Thr-Arg-Leu-Cys-Arg-Lys-Arg-Arg-Cys-Ser-Leu-Ser-Val-Ala-Gln-Glu
|
|
| LEAP-2 (38-77) (Human) / LEAP-2 (37-76) (Mouse) |
|
| 10 µCi |
|
| -20°C or lower |
|
| Each vial contains 10 µCi of lodine-125 labeled peptide. |

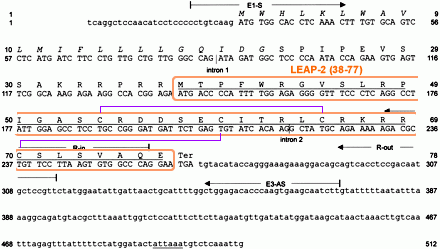 |
Human LEAP-2 precursor cDNA and deduced amino acid sequence. Coding regions are printed in capital letters. The typical secretory signal sequence 1–22 is printed in italics, whereas the positions of introns 1 (198 bp) and 2 (320 bp) of the LEAP-2 gene are marked by vertical lines. Two TATAAA sequences located 31 and 84 bases upstream of the translational start were identified. The putatively mature and antimicrobially active LEAP-2-(38–77) isolated from hemofiltrate and the putative polyadenylation signal are underlined. The position of the primers used for nested 5'-RACE-PCR and for preparative PCR from six different species is indicated. |

Alignment of LEAP-2 from mammalian species. Standard PCR carried out with primers originally designed for the human LEAP-2 gene revealed homologous LEAP-2 forms in rhesus monkey, cow, pig, mouse, and guinea pig. The depicted putative peptide sequences are deduced from the cDNA sequences obtained. Underlined amino acids represent the putative signal peptide sequences as predicted by the SignalP V2.0 program (Nielsen et al. 1997), and the mature LEAP-2 (38–77) form is hyphenated. The nucleotide sequence data reported in this paper have been submitted to the GenBank/EBI Data Bank with accession numbers AJ306405 (Homo sapiens mRNA), AJ409013 (Sus scrofa mRNA), AJ409014 (Bos taurus mRNA), AJ409054 (Cavia porcellus mRNA), AJ409055 (Mus musculus mRNA), AJ409056 (Macaca mulatta genomic DNA), AJ409063 (Mus musculus genomic DNA), AJ409064 (Homo sapiens genomic DNA), and AJ409065 (Homo sapiens alternative promoter sequence).
Abstract: Liver-expressed antimicrobial peptide 2 (LEAP-2), originally described as an antimicrobial peptide, has recently been recognized as an endogenous blocker of growth hormone secretagogue receptor 1a (GHS-R1a). GHS-R1a, also known as ghrelin receptor, is a G protein-coupled receptor (GPCR) widely distributed on the hypothalamus and pituitary gland where it exerts its major functions of regulating appetite and growth hormone (GH) secretion. The activity of GHS-R1a is controlled by two counter-regulatory endogenous ligands: Ghrelin (activation) and LEAP-2 (inhibition). Ghrelin activates GHS-R1a on the neuropeptide Y/Agouti-related protein (NPY/AgRP) neurons at the arcuate nucleus (ARC) to promote appetite, and on the pituitary somatotrophs to stimulate GH release. On the flip side, LEAP-2, acts both as an endogenous competitive antagonist of ghrelin and an inverse agonist of constitutive GHS-R1a activity. Such a biological property of LEAP-2 vigorously blocks ghrelin's effects on food intake and hormonal secretion. In circulation, LEAP-2 displays an inverse pattern as to ghrelin; it increases with food intake and obesity (positive energy balance), whereas decreases upon fasting and weight loss (negative energy balance). Thus, the LEAP-2/ghrelin molar ratio fluctuates in response to energy status and modulation of this ratio conversely influences energy intake. Inhibiting ghrelin's activity has shown beneficial effects on obesity in preclinical experiments, which sheds light on LEAP-2's anti-obesity potential. In this review, we will analyze LEAP-2's effects from a metabolic point of view with a focus on metabolic hormones(e.g., ghrelin, GH, and insulin), and discuss LEAP-2's potential as a promising therapeutic target for obesity.
Lu X, Huang L, Huang Z, Feng D, Clark RJ, Chen C. Leap-2: an emerging endogenous ghrelin receptor antagonist in the pathophysiology of obesity. Front Endocrinol. 2021;12:717544.
Abstract: The ghrelin receptor or growth hormone secretagogue receptor (GHSR) is a G-protein-coupled receptor that controls growth hormone and insulin secretion, food intake, and reward-seeking behaviors. Liver-expressed antimicrobial peptide 2 (LEAP2) was recently described as an endogenous antagonist of GHSR. Here, we present a study aimed at delineating the structural determinants required for LEAP2 activity toward GHSR. We demonstrate that the entire sequence of LEAP2 is not necessary for its actions. Indeed, the N-terminal part alone confers receptor binding and activity to LEAP2. We found that both LEAP2 and its N-terminal part behave as inverse agonists of GHSR and as competitive antagonists of ghrelin-induced inositol phosphate production and calcium mobilization. Accordingly, the N-terminal region of LEAP2 is able to inhibit ghrelin-induced food intake in mice. These data demonstrate an unexpected pharmacological activity for LEAP2 that is likely to have an important role in the control of ghrelin response under normal and pathological conditions.
M’Kadmi C, Cabral A, Barrile F, et al. N-terminal liver-expressed antimicrobial peptide 2 (Leap2) region exhibits inverse agonist activity toward the ghrelin receptor. J Med Chem. 2019;62(2):965-973
Abstract: Food homeostatic states (hunger and satiety) influence the cognitive systems regulating impulsive responses, but the direction and specific mechanisms involved in this effect remain elusive. We examined how fasting, and satiety, affect cognitive mechanisms underpinning disinhibition using a novel framework and a gamified test-battery. Thirty-four participants completed the test-battery measuring three cognitive facets of disinhibition: attentional control, information gathering and monitoring of feedback, across two experimental sessions: one after overnight fasting and another after a standardised meal. Homeostatic state was assessed using subjective self-reports and biological markers (i.e., blood-derived liver-expressed antimicrobial protein 2 (LEAP-2), insulin and leptin). We found that participants who experienced greater subjective hunger during the satiety session were more impulsive in the information gathering task; results were not confounded by changes in mood or anxiety. Homeostatic state did not significantly influence disinhibition mechanisms linked to attentional control or feedback monitoring. However, we found a significant interaction between homeostatic state and LEAP-2 on attentional control, with higher LEAP-2 associated with faster reaction times in the fasted condition only. Our findings indicate lingering hunger after eating increases impulsive behaviour via reduced information gathering. These findings identify a novel mechanism that may underpin the tendency to overeat and/or engage in broader impulsive behaviours.
Voigt K, Giddens E, Stark R, et al. The hunger games: homeostatic state-dependent fluctuations in disinhibition measured with a novel gamified test battery. Nutrients. 2021;13(6):2001.
Context: The mechanisms underlying Roux-en-Y gastric bypass (RYGB) surgery-induced weight loss and the immediate postoperative beneficial metabolic effects associated with the operation remain uncertain. Enteroendocrine cell (EEC) secretory function has been proposed as a key factor in the marked metabolic benefits from RYGB surgery.
Objective: To identify novel gut-derived peptides with therapeutic potential in obesity and/or diabetes by profiling EEC-specific molecular changes in obese patients following RYGB-induced weight loss.
Subjects and Methods: Genome-wide expression analysis was performed in isolated human small intestinal EECs obtained from 20 gut-biopsied obese subjects before and after RYGB. Targets of interest were profiled for preclinical and clinical metabolic effects.
Results: Roux-en-Y gastric bypass consistently increased expression levels of the inverse ghrelin receptor agonist, liver-expressed antimicrobial peptide 2 (LEAP2). A secreted endogenous LEAP2 fragment (LEAP238-47) demonstrated robust insulinotropic properties, stimulating insulin release in human pancreatic islets comparable to the gut hormone glucagon-like peptide-1. LEAP238-47 showed reciprocal effects on growth hormone secretagogue receptor (GHSR) activity, suggesting that the insulinotropic action of the peptide may be directly linked to attenuation of tonic GHSR activity. The fragment was infused in healthy human individuals (n = 10), but no glucoregulatory effect was observed in the chosen dose as compared to placebo.
Conclusions: Small intestinal LEAP2 expression was upregulated after RYGB. The corresponding circulating LEAP238-47 fragment demonstrated strong insulinotropic action in vitro but failed to elicit glucoregulatory effects in healthy human subjects.
Hagemann CA, Zhang C, Hansen HH, et al. Identification and metabolic profiling of a novel human gut-derived leap2 fragment. The Journal of Clinical Endocrinology & Metabolism. 2021;106(2):e966-e981.
Background/Objectives: Liver-expressed antimicrobial peptide 2 (LEAP-2) was recently identified as an endogenous non-competitive allosteric antagonist of the growth hormone secretagogue receptor 1a (GHSR1a). LEAP-2 blunts ghrelin-induced feeding and its plasma levels are modulated in response to nutritional status in humans. Despite the relevant role of ghrelin in childhood, puberty, and childhood obesity, the potential implication of LEAP-2 in these aspects remains totally unknown. We aimed to investigate the regulation of circulating plasma LEAP-2 in childhood and adolescent either lean or obese.
Methods and results: Plasma levels of LEAP-2 were analyzed in a cross-sectional study with lean and obese children and adolescents (n = 150). Circulating LEAP-2 levels were significantly higher in girls than in boys independently of whether they were obese or lean. In addition, LEAP-2 was significantly increased (p < 0.001) in pubertal than in prepubertal girls, while no changes were found in boys between both developmental stages. Moreover, in girls LEAP-2 was positively correlated with insulin, IGF-1, HOMA-IR and triglycerides and negatively with ghrelin. In boys, LEAP-2 was positively correlated with leptin and negatively with vitamin D levels.
Conclusion: This study reveals a sexual dimorphism in LEAP-2 levels in children and adolescents. These changes and the higher levels during puberty imply that LEAP-2 may contribute to some of the biological adaptations occurring during pubertal development in terms of food intake, energy balance, growth rate, and puberty onset. Future studies assessing LEAP-2 levels in longitudinal studies and its implications in growth rate, puberty onset, and reproductive hormones will help to understand the relevance of this hormone in this stage of life.
Barja-Fernández S, Lugilde J, Castelao C, et al. Circulating LEAP-2 is associated with puberty in girls. Int J Obes. 2021;45(3):502-514.
Abstract: Rheumatoid arthritis (RA) is a debilitating, chronic, inflammatory, autoimmune disease associated with cachexia. The substitutive therapy of gut hormone ghrelin has been pointed at as a potential countermeasure for the management of metabolic and inflammatory complications in RA. The recent discovery of liver-expressed antimicrobial peptide 2 (LEAP2) as an endogenous inverse agonist/antagonist of the ghrelin receptor makes feasible the development of a more rational pharmacological approach. This work aimed to assess the serum LEAP2 levels, in a cohort of RA patients, in comparison with healthy individuals and determine its correlation with inflammatory parameters. LEAP2 levels were determined by a commercial ELISA kit, plasma C-reactive protein (CRP) levels were evaluated using immunoturbidimetry, and serum levels of inflammatory mediators, namely IL-6, IL-8, IL-1β, MIP1α, MCP1, and LCN2, were measured by XMap multiplex assay. LEAP2 serum levels were significantly increased in RA patients (n = 101) compared with control subjects (n = 26). Furthermore, the LEAP2 levels significantly correlated with CRP and inflammatory cytokines, but not with BMI. These data reveal LEAP2 as a new potential RA biomarker and indicated the pharmacological control of LEAP2 levels as a novel approach for the treatment of diseases with alterations on the ghrelin levels, such as rheumatoid cachexia.
Francisco V, Tovar S, Conde J, et al. Levels of the novel endogenous antagonist of ghrelin receptor, liver-enriched antimicrobial peptide-2, in patients with rheumatoid arthritis. Nutrients. 2020;12(4):1006.
Abstract: Acyl-ghrelin administration increases food intake, body weight, and blood glucose. In contrast, mice lacking ghrelin or ghrelin receptors (GHSRs) exhibit life-threatening hypoglycemia during starvation-like conditions, but do not consistently exhibit overt metabolic phenotypes when given ad libitum food access. These results, and findings of ghrelin resistance in obese states, imply nutritional state dependence of ghrelin’s metabolic actions. Here, we hypothesized that liver-enriched antimicrobial peptide-2 (LEAP2), a recently characterized endogenous GHSR antagonist, blunts ghrelin action during obese states and postprandially. To test this hypothesis, we determined changes in plasma LEAP2 and acyl-ghrelin due to fasting, eating, obesity, Roux-en-Y gastric bypass (RYGB), vertical sleeve gastrectomy (VSG), oral glucose administration, and type 1 diabetes mellitus (T1DM) using humans and/or mice. Our results suggest that plasma LEAP2 is regulated by metabolic status: its levels increased with body mass and blood glucose and decreased with fasting, RYGB, and in postprandial states following VSG. These changes were mostly opposite of those of acyl-ghrelin. Furthermore, using electrophysiology, we showed that LEAP2 both hyperpolarizes and prevents acyl-ghrelin from activating arcuate NPY neurons. We predict that the plasma LEAP2/acyl-ghrelin molar ratio may be a key determinant modulating acyl-ghrelin activity in response to body mass, feeding status, and blood glucose.
Mani BK, Puzziferri N, He Z, et al. LEAP2 changes with body mass and food intake in humans and mice. Journal of Clinical Investigation. 2019;129(9):3909-3923.
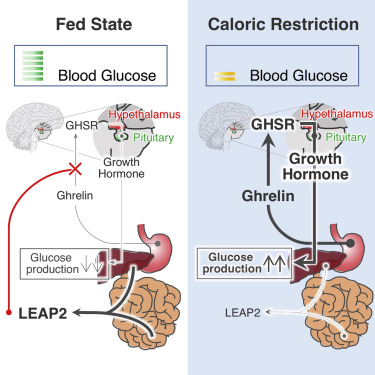
Abstract: Ghrelin, an appetite-stimulatory hormone secreted by the stomach, was discovered as a ligand for the growth hormone secretagogue receptor (GHSR). Through GHSR, ghrelin stimulates growth hormone (GH) secretion, a function that evolved to protect against starvation-induced hypoglycemia. Though the biology mediated by ghrelin has been described in great detail, regulation of ghrelin action is poorly understood. Here, we report the discovery of liver-expressed antimicrobial peptide 2 (LEAP2) as an endogenous antagonist of GHSR. LEAP2 is produced in the liver and small intestine, and its secretion is suppressed by fasting. LEAP2 fully inhibits GHSR activation by ghrelin and blocks the major effects of ghrelin in vivo, including food intake, GH release, and maintenance of viable glucose levels during chronic caloric restriction. In contrast, neutralizing antibodies that block endogenous LEAP2 function enhance ghrelin action in vivo. Our findings reveal a mechanism for fine-tuning ghrelin action in response to changing environmental conditions.
Ge X, Yang H, Bednarek MA, et al. Leap2 is an endogenous antagonist of the ghrelin receptor. Cell Metabolism. 2018;27(2):461-469.e6.
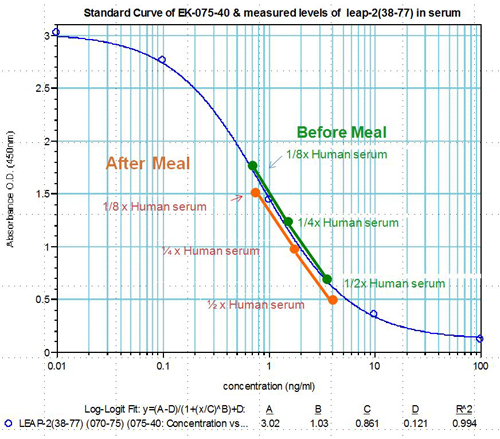
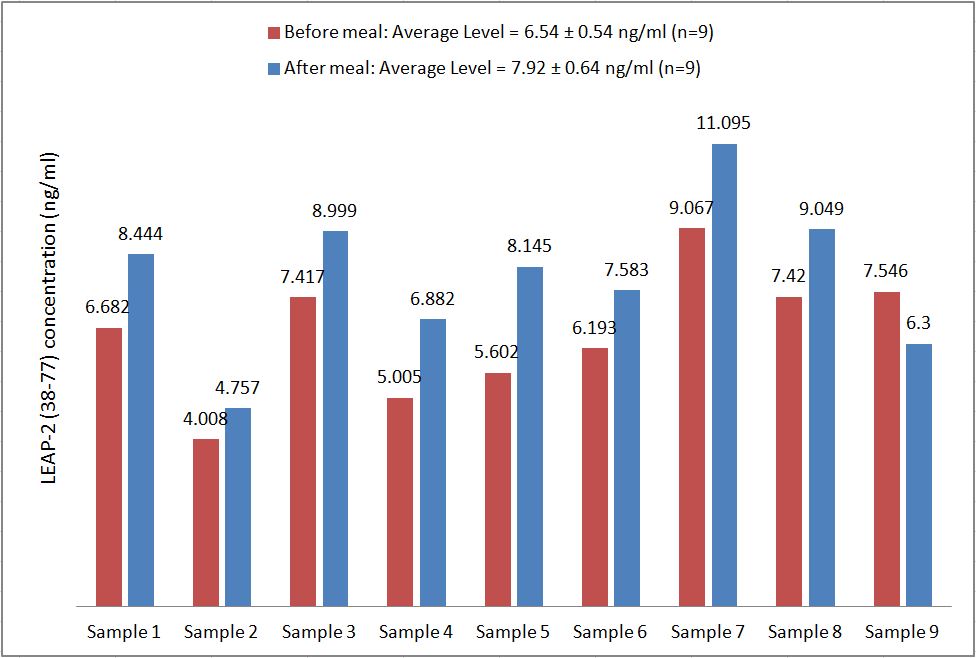
Human serum samples were assayed using our internally developed LEAP-2 (38-77) EIA kit (#EK-075-40). All samples were assayed in duplicate. On average, LEAP-2 levels increased by 21% (6.54 ng/ml vs. 7.92 ng/ml) after feeding (compared to overnight fasting).
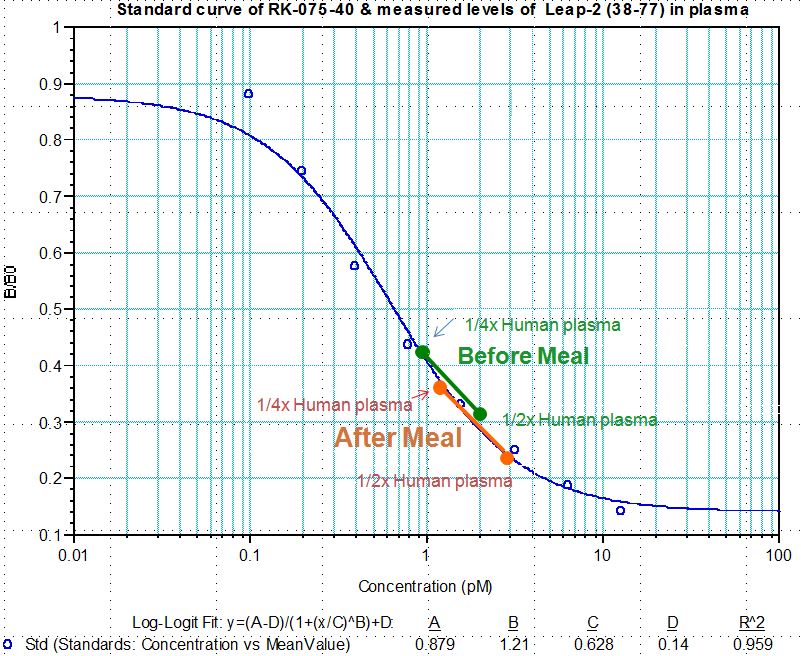
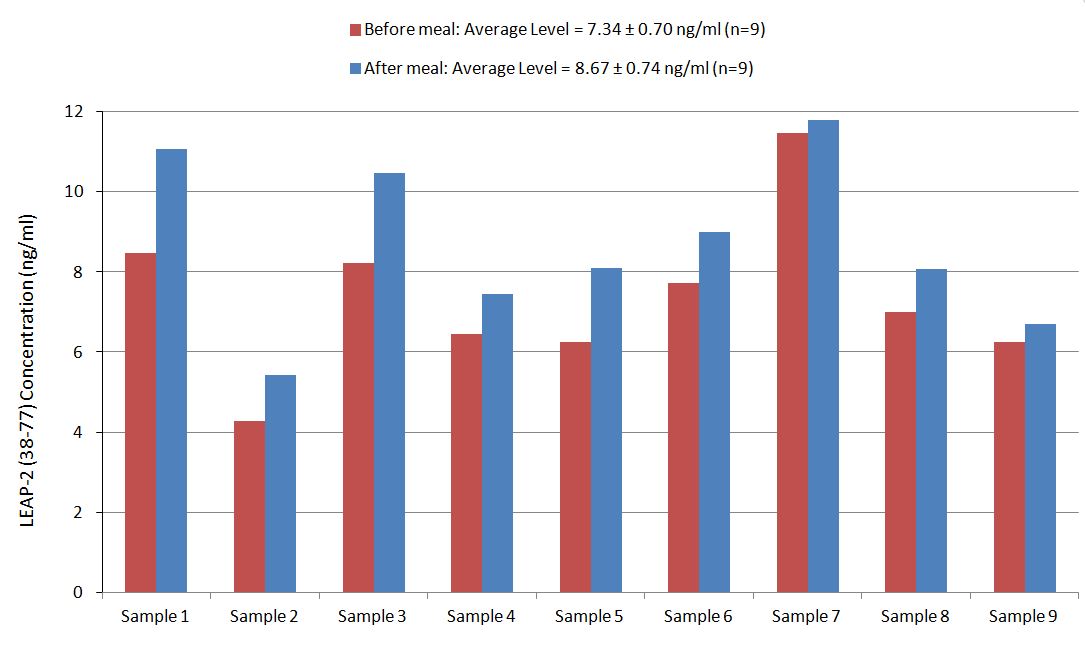
Human plasma samples were assayed using our internally developed LEAP-2 (38-77) RIA kit (#RK-075-40). All samples were assayed in duplicate. On average, LEAP-2 levels increased by 21% (7.34 ng/ml vs. 8.67 ng/ml) after feeding (compared to overnight fasting).
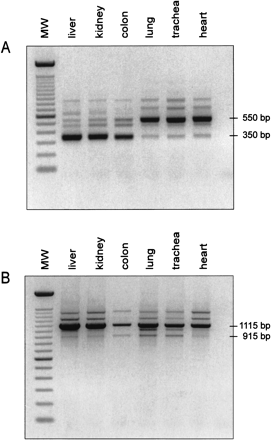 |
Alternative splicing in the LEAP-2 gene. (A) Standard PCR conducted with the primer combination E1-S and E3-AS (Fig. 2) and 20 ng of DNase-treated cDNA. LEAP-2350 is mainly expressed in liver, kidney, and colon, whereas LEAP-2550 is the main transcript in lung, trachea, and heart. PCR products at 650 bp, 720 bp, and 870 bp represent splicing variants of LEAP-2, and the band at 480 bp could be identified as a PCR artefact. (MW = marker, 100 bp DNA ladder, Life Technologies). (B) Standard PCR performed with the primers PROM-S (5'-GGTGCA GATTAGGGTGACAGTCCATC-3'), which is located 565 bp upstream of the transcriptional start depicted in Figure 2, and E3-AS. Lung, heart, and trachea exhibit the same pattern of bands, including the 565 bp shift in size caused by the upstream primer PROM-S. The main transcript identified in liver, kidney, and colon, however, changes from the completely spliced LEAP-2350 to the intron 1-retaining LEAP-2 variant. All bands obtained were characterized by DNA sequencing. |
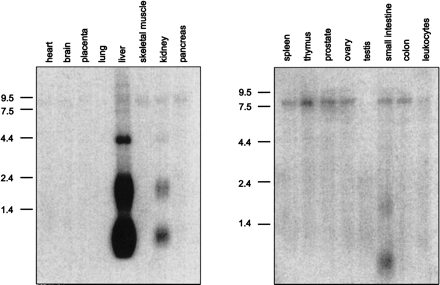 |
Northern Blot analysis. Human MTN Blots I+II (Clontech, A+B) were hybridized under high-stringency conditions with a 32P-labeled LEAP-2-specific cDNA fragment. A transcript size of 0.7 kb identified in liver, kidney, and small intestine represents the complete LEAP-2 cDNA including a poly(A)-tail of 200 bp. The signal at 2.0 kb corresponds to the LEAP-2 form coded by the distal promoter, which could also be identified in 5'-RACE-PCR, whereas the bands at 4.2 kb and 8.0 kb might either represent additional alternative promoter variants or homologous proteins related to LEAP-2. |
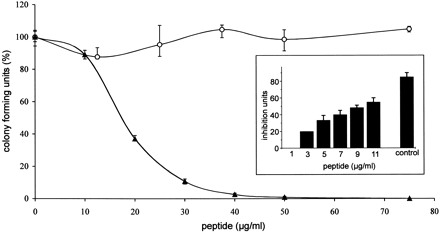 |
Antimicrobial activity of LEAP-2-(38–77) and LEAP-2-(44–77). Colony-forming unit assay of synthetic LEAP-2-(38–77) (solid triangles) and LEAP-2-(44–77) (open circles), the two main peptide forms isolated from hemofiltrate, against S. cerevisiae ATCC9763. Incubation without peptide represents 100% CFU. Synthetic and native LEAP-2-(44–77) led to similar results. The bars indicate the minimum and maximum value of the triplicates used in this representative assay. The inset depicts the dose-dependent effect of LEAP-2-(38–77) against S. cerevisiae in a radial diffusion assay. LEAP-2-(44–77) showed no effect in this sensitive antimicrobial assay. The antimicrobially active casocidin-I (11 µg/well) served as a positive control ( Zucht et al. 1998). One inhibition unit (1 IU) corresponds to 0.1 mm diameter of growth inhibition zone, and the error bars represent the S.D. calculated from three experiments performed. |
No References
| Catalog# | Product | Size | Price | Buy Now |
|---|
Social Network Confirmation